Data-Driven MRI Postprocessing
A data-driven, deep-learning based MRI postprocessing framework to improve medical image analysis
Synthetic MRI (SyMRI) enables the generation of quantitative tissue relaxation maps—T1, T2, and proton density (PD)—which can be used to synthesize a wide variety of contrast-weighted images. However, both conventional structural MRI (e.g., T1-weighted and T2-weighted scans) and current SyMRI post-processing pipelines typically rely on manually selected acquisition parameters such as repetition time (TR), echo time (TE), and inversion time (TI). These parameters are often chosen through expert-driven heuristics or visual inspection, resulting in inconsistent contrast quality across scanners and limited scalability for large cohort studies.
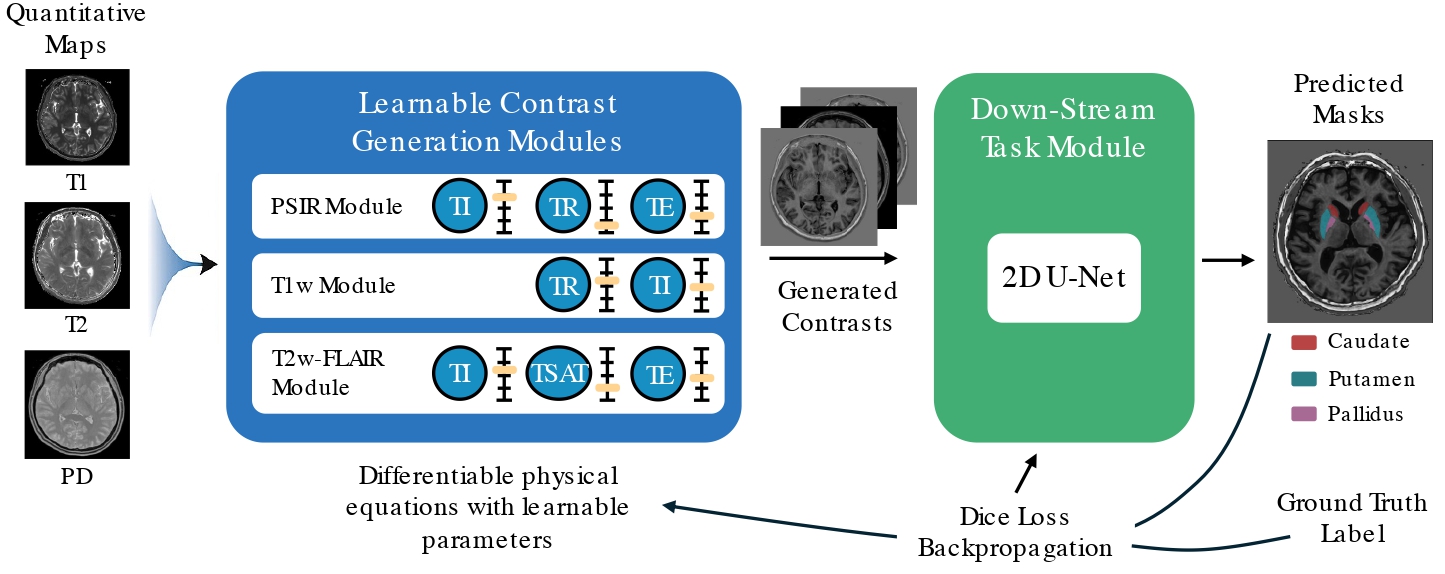
To overcome these limitations, we propose a physics-informed, learning-based framework that automatically adjusts SyMRI contrast parameters in tandem with downstream tasks such as tissue segmentation. Inspired by the concept of joint optimization of data transformations and task objectives, our approach embeds a differentiable MRI physics layer into the training pipeline. This enables end-to-end optimization of both contrast generation and segmentation modules, allowing the model to discover contrast settings that maximize task performance—while remaining faithful to underlying physical constraints.